Plant Physiology 161: 1903-1917 (2013)
Spatiotemporal seed development analysis provides insight into primary dormancy induction and evolution of the Lepidium DELAY OF GERMINATION1 Genes [W][OA]
School of Biological Sciences, Plant Molecular Science and Centre for Systems and Synthetic Biology, Royal Holloway, University of London, Egham, Surrey TW20 0EX, United Kingdom (KG, AV, GLM);
Web: 'The Seed Biology Place' - www.seedbiology.eu
University of Freiburg, Faculty of Biology, Institute for Biology II, Botany/Plant Physiology, D-79104 Freiburg, Germany (KG, AV, ABM, GLM)
Universität Osnabrück, Fachbereich Biologie, Botanik, D-49069 Osnabrück, Germany (KS, KM)
Received December 24, 2012; Accepted Feburary 19, 2013; Published Feburary 20, 2013.
DOI:10.1104/pp.112.213298
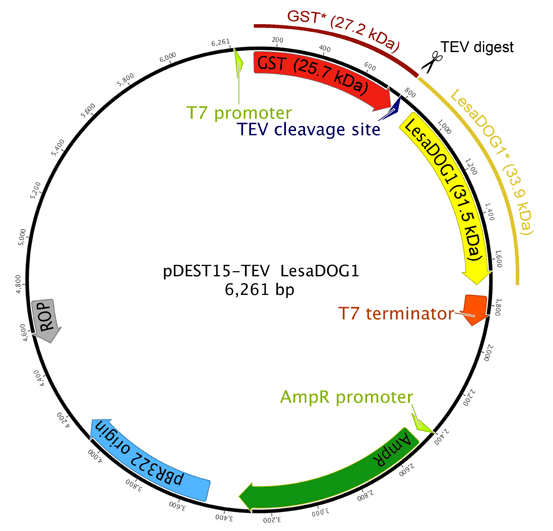
Supplemental Figure S3. Plasmid Map of modified pDEST15 vector containing a TEV cleavage site used for recombinant GST-LesaDOG1 fusion protein expression.
Predicted molecular masses of native GST and LesaDOG1 proteins are shown on the inside, predicted masses of GST* and LesaDOG1* proteins resulting from TEV cleavage of the fusion protein are shown on the outside of the plasmid map. GST-Linker(TEV)-LesaDOG1 is encoded as a continuous ORF on the plasmid. Numbers on plasmid: bp.
| Article in PDF format (2 MB) Supplementary data file (4.9 MB) |
|
|
|
The Seed Biology Place |
Webdesign Gerhard Leubner 2000 |