The Plant Cell 23: 2045-2063 (2011)
A Guideline to Family-wide Comparative State-of-the-art qRT-PCR Analysis Exemplified with a Brassicaceae Cross-species Seed Germination Case Study [W][OA]
* Both authors contributed equally to this work
The University of Nottingham, Division of Statistics, School of Mathematical Sciences, University Park, Nottingham NG7 2RD, United Kingdom (A.T.A.W.)
Received February 8, 2011; revised May 6, 2011; accepted May 27, 2011; published June 10, 2011.
www.plantcell.org/cgi/doi/10.1105/tpc.111.084103
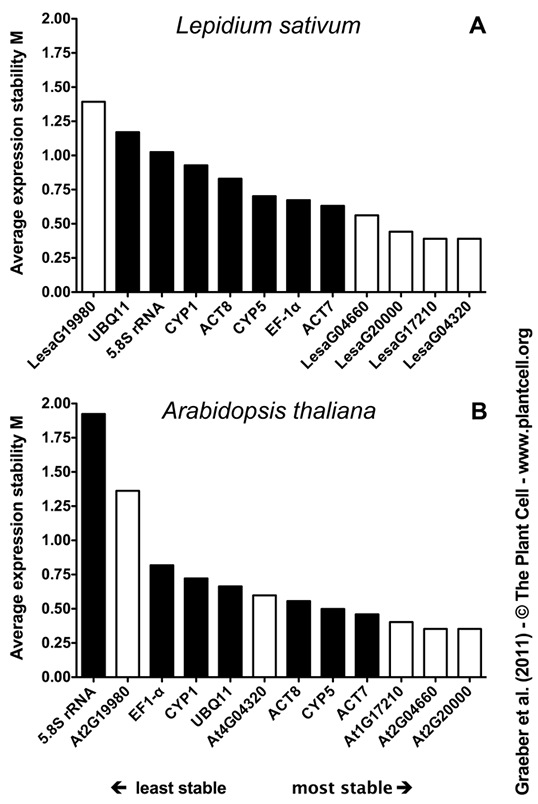 |
Figure 4. Ranking of New and Traditional Lepidium sativum and Arabidopsis Reference Genes for Seed Germination Based on their Average Expression Stability Measure M. M was determined by analysing the efficiency-corrected transcript abundances across all samples (as determined by qRT-PCR, Figure 3) via GeNORM for L. sativum (A) and Arabidopsis (B) separately. Black bars indicate traditional and white bars new reference genes which were selected from L. sativum microarray analysis (Figure 2). Note that stability measure M decreases more steadily within all the L. sativum reference genes, whereas for Arabidopsis there is a rapid decline within the three least stable genes and more stable genes do differ less among each other. A possible reason for this is the fact that due to the specific seed tissue samples the L. sativum dataset is more diverse, whereas whole-seed samples were used for Arabidopsis. The average expression stability rankings of the two species are highly similar. The orthologous pairs LesaG20000/At2G20000, LesaG04660/At2G04660 and LesaG17210/At1G17210 provide the most stable cross-species reference genes for qRT-PCR analyses of seed germination. |
Synopsis: Developmental processes like seed germination are characterised by massive transcriptome changes. This study compares seed transcriptome datasets of different Brassicaceae to identify stable expressed reference genes for cross-species qRT-PCR normalisation. A workflow is presented for improving RNA quality, qRT-PCR performance, and normalisation when analysing expression changes across species.
| Article in PDF format (1.2 MB) Supplemental data file (156 KB) |
|
|
|
The Seed Biology Place |
Webdesign Gerhard Leubner 2000 |